Message from the Chair
Our department has a rich history of excellence in research, education and training.
Education
Explore the broad field of biochemistry with our BS and BA undergraduate programs. Or advance your career with our PhD and MS programs, as well as dual-degree programs that combine an MS in Biochemistry with JD, MBA, or MD degrees.

Who We Are
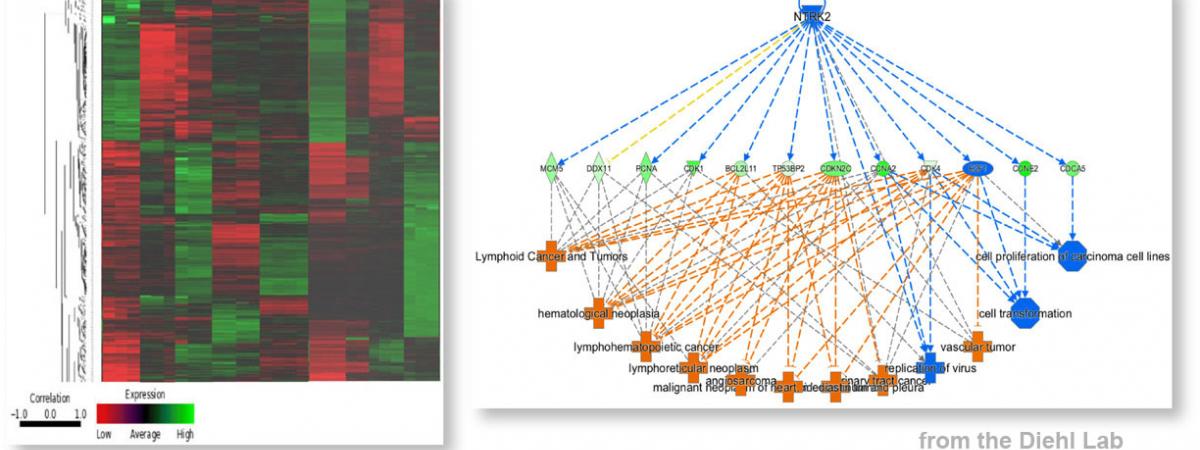
Our faculty members are leaders in a wide range of biomedical research areas.
News
J. Alan Diehl receives the Case Medal for Excellence in Health Science Innovation
October 15, 2025When J. Alan Diehl arrived at Case Western Reserve University in 2019, the Department of Biochemistry in the School of Medicine was at a crossroads—ready for a…Wood Prize for Biochemistry MS
August 06, 2025Congratulations to Clare Mewhinney, our 2025 winner of the Wood Prize for excellence in the Biochemistry MS program!Biochemistry senior prizes and honors
April 30, 2025Congratulations to all of our graduating Biochemistry seniors, who presented very impressive capstone posters this week! We also celebrate these seniors for…